Association of AXIN1 With Parkinson’s Disease in a Taiwanese Population
Article information
Abstract
Objective
A meta-analysis of locus-based genome-wide association studies recently identified a relationship between AXIN1 and Parkinson’s disease (PD). Few studies of Asian populations, however, have reported such a genetic association. The influences of rs13337493, rs758033, and rs2361988, three PD-associated genetic variants of AXIN1, were investigated in the present study because AXIN1 is related to Wnt/β-catenin signaling.
Methods
A total of 2,418 individuals were enrolled in our Taiwanese cohort for analysis of the genotypic and allelic frequency. Polymerase chain reaction–restriction fragment length polymorphism analysis was employed for rs13337493 genotyping, and the Agena MassARRAY platform (Agena Bioscience, San Diego, CA, USA) was used for rs758033 and rs2361988 genotyping in 672 patients with PD and 392 controls. Taiwan Biobank data of another 1,354 healthy controls were subjected to whole-genome sequencing performed using Illumina platforms at approximately 30× average depth.
Results
Our results revealed that rs758033 {odds ratios [OR] (95% confidence interval [CI]) = 0.267 [0.064, 0.795], p = 0.014} was associated with the risk of PD, and there was a trend toward a protective effect of rs2361988 (OR [95% CI] = 0.296 [0.071, 0.884], p = 0.026) under the recessive model. The TT genotype of rs758033 (OR [95% CI] = 0.271 [0.065, 0.805], p = 0.015) and the CC genotype of rs2361988 (OR [95% CI] = 0.305 [0.073, 0.913], p = 0.031) were less common in the PD group than in the non-PD group.
Conclusion
Our findings indicate that the rs758033 and rs2361988 polymorphisms of AXIN1 may affect the risk of PD in the Taiwanese population.
After Alzheimer’s disease (AD), Parkinson’s disease (PD) is the most prevalent neurodegenerative disorder [1]. Evidence increasingly shows that genetic factors play major roles in PD. However, few of the heritable components have been identified [2]. Pathologically, the hallmarks of PD are abnormal α-synuclein aggregation and dopaminergic neuron loss in the substantia nigra (SN) [3]; these factors lead to tremors, rigidity, bradykinesia, and stooped posture, among other major symptoms of PD [4]. Although the detailed cause of dopaminergic neuron loss remains unclear, accumulating evidence indicates the important role of neuroinflammation in neurodegenerative diseases such as PD [5]. In one study, activated astrocytes and microglia were used for chronic release of proinflammatory cytokines, contributing to the degeneration of dopaminergic neurons in the SN [6].
Several signaling pathways, such as that of Wnt, are believed to be involved in neuroinflammation in AD and PD [7-9]. The Wnt signaling pathway is commonly considered to have canonical and noncanonical components, with the canonical pathway believed to promote anti-inflammatory activity [7]. AXIN1 promotes the GSK-3β phosphorylation of β-catenin and subsequent degradation of β-catenin in the canonical Wnt pathway, resulting in reduced Wnt expression [10]. A study also discovered a regulatory role of the canonical Wnt pathway in neuroinflammation [11]. Because AXIN1 negatively regulates this pathway, it may affect the pathogenesis of PD by promoting brain inflammation and impairing adult neurogenesis [8,12].
Recently, the relationship between AXIN1 and PD was reported in a meta-analysis of locus-based genome-wide association studies investigating genomic convergence and linking PD with inflammation [12]. AXIN1 expression was upregulated in the SN of patients with PD, and its expression in the SN was discovered to be significantly modulated by the intronic single nucleotide polymorphism (SNP) rs13337493 [12]. Additionally, rs13337493 is in linkage disequilibrium with rs758033 and rs2361988. Data on quantitative trait loci revealed that in the basal ganglia, AXIN1 expression was modulated by the SNPs rs13337493, rs758033, and rs2361988 [12]. The present research is a case-control study in which the genotypic and allelic frequencies of AXIN1 rs13337493, rs758033, and rs2361988 were examined in 672 and 1,746 Taiwanese individuals with and without PD, respectively, to determine whether these novel genetic loci are relevant to the incidence of PD among the population of Taiwan.
MATERIALS & METHODS
Ethics statement
The Institutional Review Board of Chang Gung Memorial Hospital (CGMH; ethical license No: 201701458B0C601 and 201800893B0C102) approved the protocol employed in this study, which was conducted in accordance with the 1975 (revised in 2013) Declaration of Helsinki. Every participant provided informed consent in writing.
Patient population
In total, 2,418 Taiwanese participants were included in the study. Neurological clinics at CGMH were the source of the 672 patients with PD. PD was diagnosed by an experienced movement disorder specialist (Yih-Ru Wu) using the UK Parkinson’s Disease Society Brain Bank criteria [13].
Additionally, 392 healthy controls were recruited from the same neurological clinics. We also included Taiwan Biobank data on 1,354 additional healthy controls.
Genetic analysis
The three SNPs investigated, which may modulate AXIN1 expression in the SN, were rs13337493, rs758033, and rs2361988.
Polymerase chain reaction (PCR)–restriction fragment length polymorphism analysis was used to confirm rs13337493. The following primer sequences were used: 5'-TGCGGCATATTGTGTCCTAA-3' (forward primer) and 5'-GCGACAAGAGGACAAGAGTG-3' (reverse primer). The Hpy166II (New England Biolabs, Ipswich, MA, USA) enzyme recognition site was GTNNAC (the polymorphic site is underlined). The material for and process of PRC amplification are described in Supplementary Material 1 (in the online-only Data Supplement). Hpy166II was employed to digest the amplified PCR fragments, after which they were separated on a 1.0% agarose gel (allele A, 454-bp fragment; allele G, 274- and 180-bp fragments).
The Agena MassARRAY platform with iPLEX Gold chemistry (Agena Bioscience, San Diego, CA, USA) was employed to genotype rs758033 and rs2361988. Assay Designer version 4.0 was used (in accordance with the manufacturer’s guide) to determine the specific PCR primer and extension primer sequences (Supplementary Table 1 in the online-only Data Supplement). Multiplex PCR was conducted and is described in Supplementary Material 2 (in the online-only Data Supplement). Residual salt was removed by adding a cation exchange resin. Subsequently, the matrix pad from a SpectroCHIP array (Agena Bioscience, San Diego, CA, USA) was loaded with 7 nL of the purified primer extension reaction mix. A MassARRAY Analyzer 4 (Agena Bioscience) was employed for SpectroCHIP evaluation, and TYPER 4.0 software (Sequenom, San Diego, CA, USA) was employed to perform clustering analysis of the calling.
Statistical analysis
We evaluated the Hardy–Weinberg equilibrium of the genotypic frequencies in the PD and non-PD groups. The allelic and genotypic distributions of the two groups were compared with the chi-square test. For analysis of the PD–SNP associations, odds ratios (ORs) with 95% confidence intervals (CIs) were calculated. As this study involved three independent genetic loci, we made a modest correction using the Bonferroni method for multiple comparisons with statistical significance defined at p < 0.017. The power of our study was calculated by using G*power 3.1 software.
RESULTS
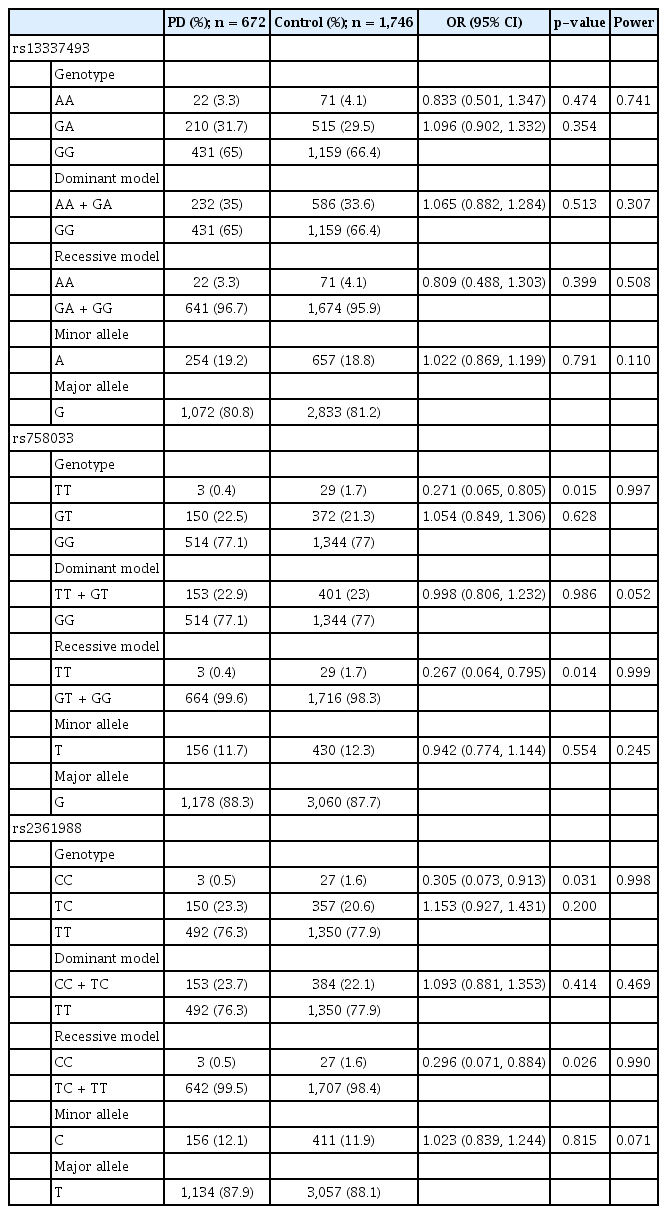
This study recruited 2,418 individuals: 672 with PD and 1,746 controls. The genotypic frequency distributions of three SNPs in the two groups were compared (Table 1). The rs758033 TT genotype was less frequent in the PD patients than the controls (OR [95% CI] = 0.271 [0.065, 0.805], p = 0.015) and had a lower risk of PD compared with that of the genotypes GT + GG in the recessive model (OR [95% CI] = 0.267 [0.064, 0.795], p = 0.014). The rs2361988 CC genotype was also less frequent in the patients with PD than the controls (OR [95% CI] = 0.305 [0.073, 0.913], p = 0.031), and in the recessive model, the individuals with this CC genotype had a lower risk of PD than did those with the genotypes TC + TT [OR (95% CI) = 0.296 (0.071, 0.884), p =0.026]. The two groups had similar rs13337493 genotypic frequencies.
DISCUSSION
The associations between three AXIN1 SNPs (rs13337493, rs758033, and rs2361988) and susceptibility to PD were investigated in the present study. Our results revealed that the AXIN1 SNP rs758033 was associated with the risk of PD under the recessive model, and there was a trend toward a protective effect of rs2361988.
AXIN1, known as a tumor suppressor gene, inhibits Wnt signaling and is implicated in several cancers, such as colorectal adenocarcinoma [14], non-small-cell lung carcinoma [15], breast cancer [16], hepatocellular carcinomas [17], medulloblastomas [18], and melanoma [19]. Melanoma has also been found to have a positive association with PD [20,21]. which may be due to LRRK2 mutation and its linkage in the Wnt signaling pathway [22,23].
Vertebrates have two alternative AXIN splicing products, AXIN1 and AXIN2, which are almost functionally equivalent in the canonical Wnt pathway [24,25]. There are two types of Wnt signaling pathways: canonical and noncanonical Wnt signaling pathways. Canonical Wnt signaling promotes anti-inflammatory activity, whereas noncanonical Wnt signaling serves as a proinflammatory mechanism [7].
Binding of Wnt ligands to Frizzled family receptors and lowdensity lipoprotein receptor-related protein 5/6 leads to activation of the scaffold protein Disheveled and AXIN1 recruitment, further resulting in disassembly of the β-catenin destruction complex. Disrupting this complex—originally composed of adenomatous polyposis coli, AXIN1, GSK3β, and CK1—impairs the degradation and phosphorylation of β-catenin. Thereafter, this protein accumulates in the cytoplasm and is subsequently translocated to nuclei, where it binds to the transcription factor T-cell factor/lymphoid enhancer factor, in turn inducing Wnt gene expression [24,26,27]. This pathway may eventually promote anti-inflammatory activity [7]. Impaired canonical Wnt/β-catenin signaling was found to predispose midbrain dopaminergic neurons to death in PD and in other major neurodegenerative disorders [28]. In a murine model of 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine, neuroprotective and neurorestorative effects in the PD-injured brain were activated by canonical Wnt/β-catenin signaling [28].
Although the direct mechanism through which AXIN1 is related to PD is uncertain, various studies have implicated the dysregulation of Wnt/β-catenin signaling in the pathogenesis of neuroinflammatory and neurodegenerative disorders such as AD [7,29] and PD [9]. Furthermore, Song et al. [25] reported that AXIN1 crucially affects the regulation of Wnt/β-catenin signaling. AXIN1 gene expression was recently found to be upregulated in a murine PD model [8]. In addition, knockdown of AXIN2, a functionally interchangeable isoform of AXIN1, was reported to increase Wnt/β-catenin signaling pathway activity and thus enhance mitochondrial biogenesis and dopaminergic neurogenesis in rats with PD [30].
AXIN1 was identified in the gene list of the top pathways related to neurodegeneration in the study by Foo et al. [31], but it was not proven to be significant. Thus, the three single nucleotide variants (SNVs) were not analyzed in this genome-wide association study of a large Han Chinese cohort. The significant association between the two SNVs (rs758033 and rs2361988) and PD could be attributed to their altered gene expression level of AXIN1, which has been revealed in Mohammad Saeed’s study [12].
We revealed for the first time that in a Taiwanese population, the recessive model and TT genotype of AXIN1 rs758033 were associated with lower risks of developing PD, and the recessive model and CC genotype of rs2601988 tended to be associated with a lower risk of PD. More case-control studies are needed to confirm the AXIN1—PD association and the associations of AXIN1 with other neurodegenerative diseases; such research could aid in the design of novel therapies.
Supplementary Materials
The online-only Data Supplement is available with this article at https://doi.org/10.14802/jmd.21073.
SUPPLEMENTARY MATERIAL 1
PCR amplication of rs13337493 (restriction fragment length polymorphism analysis)
SUPPLEMENTARY MATERIAL 2
Multiplex PCR of rs758033 and rs2361988 (Agena MassARRAY platform; Agena Bioscience, San Diego, CA, USA)
Supplementary Table 1.
Sequences of primers used for genotyping in this study
Notes
Conflicts of Interest
The authors have no financial conflicts of interest.
Funding Statement
This work was supported by Minister 203 of Science and Technology, Taiwan (MOST 107-2314-B-182A-045-MY2) and Chang Gung Memorial Hospital, Taipei, Taiwan (CMRPG3H0313).
Author Contributions
Conceptualization: Yih-Ru Wu. Data curation: Hwa-Shin Fang, Yih-Ru Wu. Formal analysis: Chih-Ying Chao, Hwa-Shin Fang, Wen-Lang Fan, Chun-Chieh Wang. Funding acquisition: Yih-Ru Wu. Investigation: Yih-Ru Wu. Methodology: Hon-Chung Fung, Yih-Ru Wu. Project administration: Yih-Ru Wu. Resources: Hon-Chung Fung, Yih-Ru Wu, Po-Jung Huang, Wen-Lang Fan. Software: Wen-Lang Fan. Supervision: Wen-Lang Fan, Yih-Ru Wu. Validation: Chun-Chieh Wang. Visualization: Chih-Ying Chao, Chun-Chieh Wang. Writing—original draft: Hwa-Shin Fang. Writing—review & editing: Yih-Ru Wu.
Acknowledgements
We thank all the patients and the staffs at the Department of Neurology of the Chang Gung Memorial Hospital, Linkou Medical Center, Genomic Medicine Core 200 Laboratory of Chang Gung Memorial Hospital, Linkou Medical Center and Whole-Genome Research Core Laboratory of Human Diseases, Chang Gung Memorial Hospital, Linkou Medical Center, Keelung, Taiwan, for their valuable support of this study.